Unlock the Microbiome with Metagenomics
The only sequencing method that identifies all the microbes in a sample by analysing entire genomes.
Sequencing DNA
Metagenomics works by sequencing DNA — the genetic blueprint found in every living organism. By extracting and sequencing all DNA in a sample, metagenomics captures the full biological content of the environment being analysed.
DNA Measured in Bases
DNA is made up of four bases (A, T, C, and G). Metagenomic sequencing reads billions of these bases, creating detailed genetic data that reflects the true complexity of microbial communities.
Detecting All Microbes
Because metagenomics sequences all DNA, it can detect every microbe present — including bacteria, viruses, fungi, parasites, and archaea. This includes unknown or unexpected microbes, without needing prior assumptions or targeted primers.
Aquaculture Relevance
Our metagenomics reports show which microbes are present, their relative abundance, and their relevance to aquaculture. This transforms raw genetic data into actionable insights for health management, water quality, and production performance.
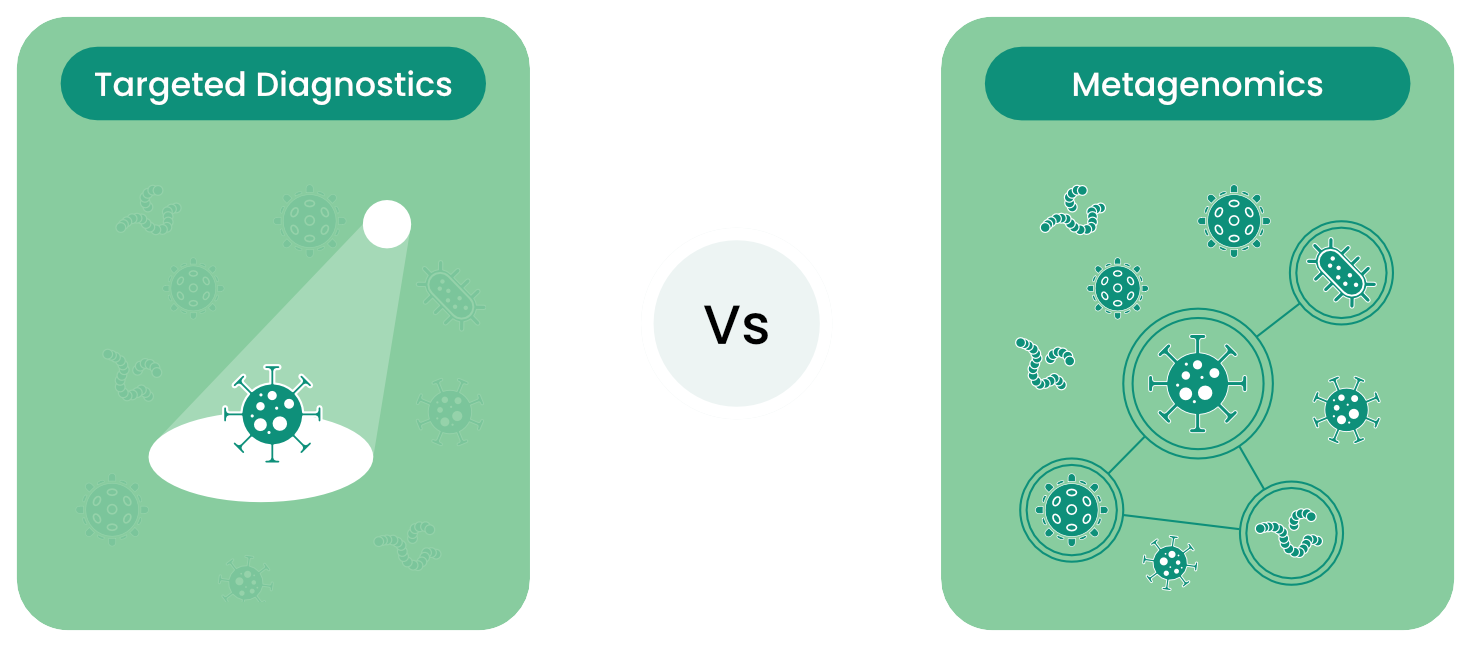
When You Need the Full Picture
Traditional diagnostics are like torches: they show only what you choose to point them at. Metagenomics is a floodlight, illuminating the entire microbial landscape in one go.
By removing targeting bias, untargeted detection reveals:
- Emerging or unexpected pathogens
- Opportunistic microbes that indicate imbalance
- Beneficial microbes that help maintain stability
The Metagenomics Workflow
1. Sample Collection
Water, mucus, or biofilm samples are collected to represent the microbiome interacting with livestock.
2. Sequencing
All DNA in the sample is extracted and sequenced without amplification ensuring an unbiased dataset.
3. Bioinformatics Analysis
Sequenced DNA is cross referenced against our genomic database to identify microbes and reveal their abundance.
4. Actionable Insights
Results are translated into clear microbiome insights, highlighting pathogens, beneficial microbes, and informing you of their relevance to aquaculture.
Application in Aquaculture
Disease Prevention
Detect pathogens early, including emerging and low-abundant threats, before they escalate into clinical disease.
Water Quality Management
Understand the microbial processes influencing nitrogen cycling, organic load, and system stability.
Monitor Microbes
Establish a healthy microbial baseline unique to your farm and monoitor changes to detect early signs of imbalance.
Optimise Operations
Track how management decisions, environmental changes, and biosecurity measures influence microbial dynamics across production systems.
Technology Comparison
Whole genome metagenomics (WGM) stands apart from other microbial detection methods by enabling both species and strain level identification of expected and unexpected microbes. In one sample metagenomics reveals the presence and abundance of bacteria, viruses, fungi, parasite and archaea.
Ready to start your microbiome journey?
Discover how total microbiome analysis can work on your farm.
Contact Us